Explainer
- Explainer
- Coronavirus pandemic
The same coronavirus strain turned up in an Australian and a tiger. How?
Tracking a virus's “genetic passport” has revolutionised how we fight pandemics. What does genomic testing tell us about COVID-19's journey around the world – and the latest outbreaks in Australia?
By Sherryn Groch and Liam Mannix
Nadia had a cough and it wouldn’t go away. She didn’t feel like eating. X-rays and ultrasounds, blood work, all turned up nothing. So a decision was made to test Nadia – a four-year-old Malayan tiger at the Bronx Zoo in New York – for the latest bug going around, the one that had already put much of the world, including her zoo, into lockdown: COVID-19.
On the other side of the globe, a 42-year-old woman in Melbourne had come down with similar symptoms to Nadia – the same dry cough, the fatigue. And, when the tests came back from the labs, the sample of the virus found in both human and tiger was near identical too. Within days, another swab with that tell-tale genetic signature would be extracted from a 21-year-old in Taiwan.
How does the same strand of virus end up in a woman in Melbourne, someone in Taiwan, and a tiger in the Bronx?
Unlike historic plagues, COVID-19 is not spreading in a straight line, from house to house or country to country. Instead, versions of the virus – each with their own tiny genetic tweaks – are jumping from continent to continent via plane (and cruise ship), and crossing back on each other. The story scientists who specialise in reading COVID’s genes see is that of the world’s first truly globalised pandemic.
Since the genetic code of the virus was first extracted from a sample in Wuhan, China, where the outbreak began, and published, hundreds of labs around the world have followed suit. Their efforts to join the dots among patients, under the microscope as well as on the ground, mean this pandemic is being tracked like never before – in real time. Already the family tree of COVID-19 is helping us understand where the outbreak came from, how it took hold in different continents – and where it’s going next.
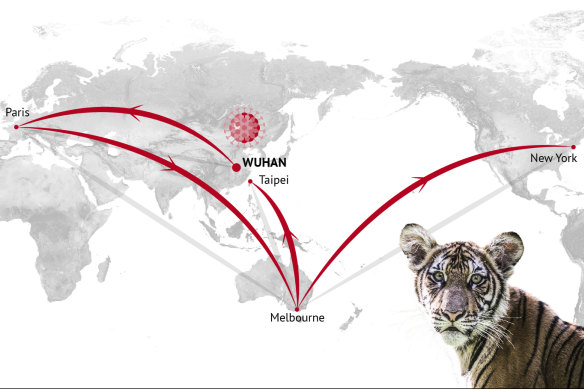
DNA detectives have tracked one distinct strand of coronavirus from Wuhan through Europe, Australia and Taiwan all the way to Nadia the tiger, pictured, at the Bronx Zoo in New York.Credit: Stephen Kiprillis, Kirsten Burghard
There are more viruses in the world than stars in the sky. They are found in almost every corner of the Earth, they fill its oceans. If lined up, side by side, scientists say they would fill our galaxy too. Most of them can’t infect us and the few that can we catch mostly from animals – in that knife-edge moment of chance when a pathogen long circulating in one species leaps to another.
Ten thousand times smaller than a grain of sand, viruses can drive forward evolution and unleash hell – think of them as zombie shells of protein waiting to spring into action.
“They’re half living, half dead,” says Professor Seshadri Vasan, from behind several Get Smart-style levels of security at the CSIRO’s dangerous pathogens lab.
Unlike bacteria, which can thrive wherever it's warm and wet, viruses need a host to survive. They are the pirates of the microscopic world, commandeering the cellular machinery of other organisms to make copies of themselves – and spread.
All they need is a way in.
“That’s why it’s Mission Impossible here at the lab,” Vasan says.
Some time in late 2019, a coronavirus now known as SARS-CoV-2 found a door into the human genome – this tiny ball of protein evolved a “spike” able to unlock our cells by binding to ACE2 receptors, a type of enzyme found throughout our bodies. In the bustling city of Wuhan, the virus jumped from an animal into a person, most likely through the handling and slaughter of wildlife for the meat, fur and traditional medicine trade. From there, the germ spread like wildfire between humans – infecting more than six million people and killing 400,000 within its first six months.
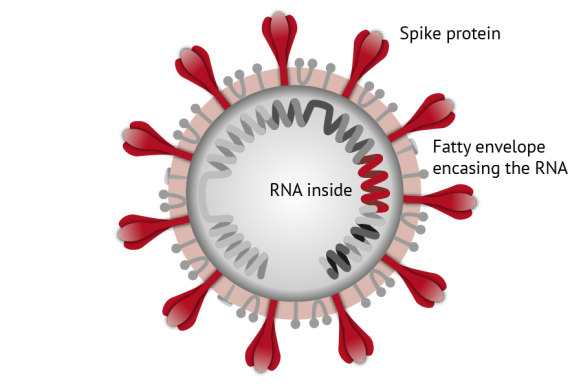
"Trouble wrapped in protein": Inside its spiky outer shell the coronavirus contains a folded-up strand of RNA.Credit: Kirsten Burghard
Beneath the coronavirus’s signature spikes, you’re looking at an envelope of fat encasing a long strand of RNA. Like our own double-stranded DNA, this holds the virus’s instruction manual for making more copies of itself.
“It’s like a manuscript written in a language made up of only four letters: A, C, G and U, which is more commonly called T,” Vasan says. How they are arranged on the page can change how the virus behaves.
SARS-COV-2’s manual is long – made up of 30,000 letters or nucleotides. By contrast, the influenza strain that caused the world’s last pandemic in 2009, swine flu, was just 13,500 letters long, Vasan says. “This is more than twice the size, it’s a very big virus.”
It is also more deadly.
Once a virus is inside a cell, it sets up shop, replicating at speed. Imagine photocopying this viral manuscript, Vasan says but each copy is then used to make the next. Before too long, mistakes – smudges or missing letters – will start to creep in. These typos are known as mutations. Some of them slow the virus and are dropped from the text over time. Most are silent – they don’t affect the virus’s structure or function – and so you can still understand the sentences. But then there are the rarer mistakes, the ones that improve the virus.
“Perhaps it's a really bad manuscript,” laughs Professor Francois Balloux, who heads up the genetics institute at University College London.
Before it emerged in humans, scientists say two key mutations had already primed SARS-CoV-2 to jump species lines and spread further than the other two dangerous coronaviruses to come before it, SARS-CoV-1 and MERS-CoV. One change – allowing the virus to fuse cells together and tunnel its way through the body faster – came down to just three letter changes in its RNA code, says Dr Denis Bauer at the CSIRO.
But now that it is circulating in humans, scientists say the coronavirus is mutating at a fairly “stable” rate – about 20 mutations a year, Balloux calculates. That’s a little faster than he expected, although not as fast as less predictable viruses such as influenza, which requires an updated vaccine every season to keep up with all its mutant stains.
Runaway COVID-19 outbreaks in parts of Europe and the Americas have fuelled speculation that more deadly or infectious strains of the virus may be circulating already – some early studies have suggested as many as 30 kinds of SARS-CoV-2. But most scientists stress the mutations so far haven’t changed how the virus behaves.
Nadia’s particular virus, for example, when examined under the microscope by Dr Leyi Wang at the University of Illinois, had racked up just six recurrent mutations or protein changes since it travelled out of Wuhan to her zoo in New York.
Critically, Vasan notes, mutations in the RNA code or even observed in how the virus behaves in the petri dish must be matched against clinical outcomes in patients to truly warrant the distinction. “Otherwise they might look like they’re having an effect when they’re not,” he says. “I’m not seeing anything sinister in the mutations so far.”
At the Bedford lab, which pulls together global genomic data on COVID-19 through the platform Nextstrain, Dr James Hadfield agrees. Most of the time when a virus starts to behave differently it’s down to its environment not its biology – as it encounters different healthcare systems, population vulnerabilities and public health interventions.
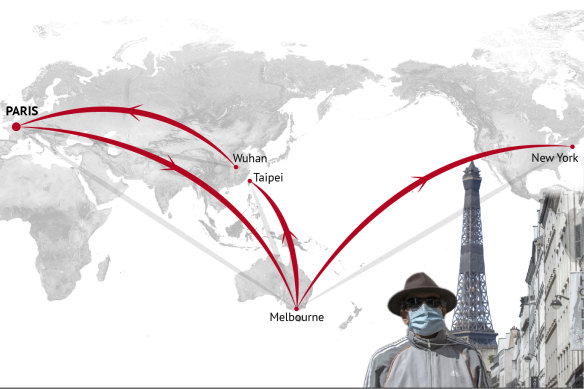
Nadia's virus was traced back through Europe to the original outbreak in China.Credit: Stephen Kiprillis, Kirsten Burghard
But mutations do come in handy for tracking the virus. Over time, differences between samples (good, bad and not worth mentioning) build up, leaving tiny molecular clues for scientists to follow – each genome is timestamped to when (and where) it was collected.
The small team behind Nexstrain works around the clock to crunch this data – when Hadfield signs off in New Zealand, Dr Emma Hodcroft and her colleagues in Switzerland are ready to take over, followed by Dr Trevor Bedford’s team in Seattle.
Hodcroft says the very first genomes out of China were just one mutation apart. Six months on, the branches of the tree still aren’t very long – samples of the virus vary by only about 10 recurrent mutations on average and their roots all go back to a common relative: that first genome sequenced in Wuhan. That’s how scientists know the spillover from animals into humans happened recently in China, likely in November 2019.
Evolutionary biologist Professor Edward Holmes says even if there were earlier lineages of the virus that died out before they could be detected (and sequenced), the jump to humans from animals still happened recently.
“Now we’ve moved from the sparks being thrown off by the Chinese epidemic, to individual epidemics starting in certain countries,” Hodcroft says.
Local mutations may develop as the virus spreads in a specific community, particularly with much of global travel grounded, but no country has a distinct national strain yet. “There is no US strain or Australian strain,” Balloux says.
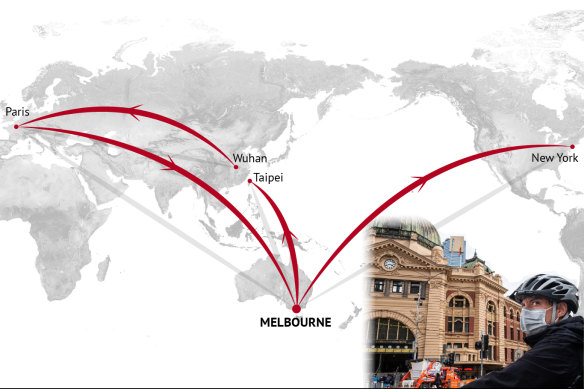
On April 1, a woman was swabbed in Melbourne. Within days, virtually the same genetic signature would be extracted from Nadia in New York.Credit: Stephen Kiprillis, Kirsten Burghard
Update: Tracking the genomics of clusters in Victoria and NSW
July 16 - Genomic analysis of the latest surge in Victorian infections has traced many cases back to just one or two sources – an outbreak among private security guards hired to man the state's hotel quarantine sites. This new surge of cases is distinct genetically from Australia's first peak in March, officials say, suggesting it broke through infection controls at just one or two sites and then spawned larger family clusters across Melbourne's north. The Victorian government says genomic investigation was behind its move to lock down 12 known hotspot suburbs – and then on July 8 metropolitan Melbourne itself – in a bid to beat back the growing tide of infections. But the data will not be released publicly until a judicial inquiry into failures within the hotel quarantine system gets underway. Over the border in NSW, a number of fresh outbreaks, including at a pub, have since been traced via genetics back to recent Melbourne cases. Read on to find out how.
In Australia, COVID-19 cases are comparatively few but our virus is among the most sequenced in the world. More than 60 per cent of all confirmed Victorian cases have now had their sample sequenced, Dr Sebastian Duchene of the Peter Doherty institute for Infection and Immunity says. And in NSW, genomic detectives have tracked early cases in Australia back to Iran as well as China, based on distinct mutations.
When you compare Australia’s viral tree with somewhere like the US, you see more genetic diversity, Hadfield says – most cases tend to be imported from overseas, giving us not just one initial case or “patient zero” but hundreds. But where clusters emerge without a clear source, genomic data can help contact tracers crack the case.
Sequencing or tracking the virus’s “genetic passport” has now revolutionised how health authorities fight pandemics, says Professor Benjamin Howden at the Doherty Institute, and will become “critically important as countries ease restrictions”. In some cases in Victoria, he says, clusters in which people had no known links to each other were uncovered after the lab team traced a connection in the virus’s genes.
“Usually, when you do contact tracing, everyone in a cluster has the same virus,” Duchene says.
This observation is what makes Nadia and her “Melbourne” strain so strange, on the face of things.
Every dot below is a Victorian case, and every line is a link between that patient and another made by contact tracers. Dots of the same colour share the same genetic virus signature as others. Some clusters, such as the large 75-person outbreak that erupted in March (shown in red down the bottom) were also linked genetically to other clusters – although no connections were found on the ground.
Victorian clusters between January 25 and April 29 were linked by genetics as well as contact tracing. Credit: Source: The Doherty Institute
Still, the world’s genomic dataset on COVID-19 is both museum and live recording – it’s not over yet.
Scientists stress they have only part of the picture – most virus lineages infecting people will not be sequenced. Even that first sample published in Wuhan did not come from a definitive “patient zero” but one of the earliest known cases: a 41-year-old worker in the wet market where the virus is suspected to have first broken out or circulated.
“We might not ever find the very first because so many people get no symptoms or mild symptoms,” Balloux says.
Vasan warns there’s a danger in any outbreak: in the race to find a cure, scientists could pick up the first sample that comes their way and run with it - in the wrong direction. When the Doherty Institute became the first lab outside China to sequence and grow the virus, it was passed like a baton in a relay, straight to Vasan at the CSIRO for mass production as his team test some of the world’s top vaccine candidates.
“But in the back of my mind, I wanted to be sure what we were working with was a typical strain,” Vasan says.
So, he called in experts in artificial intelligence, including Bauer, to help screen samples for unusual mutations, particularly in areas where changes could cause a real difference to how the virus behaves, such as in its “unstable” and much-discussed spike. The results were “reassuring” – they found that the first Wuhan strain, now used all around the world as a reference, was a good sample, as were the baseline strains used in Washington, Sydney, Melbourne and elsewhere. “Even labs in the UK use our first Victorian strain now,” Vasan says.
But they did find some outliers, including a NSW sample which contained a curious deletion. Another, collected in Italy, had mutations in common with both a Sydney virus cluster and one on the other side of the world in Chongqing, China.
“We’re not sure why,” Bauer says. “It could have been a recombination, where someone is infected with two lineages of virus at the same time that gave rise to this particular version.”
The probability of two different viral lineages evolving the exact same mutations on their own is very low, Bauer says, though Vasan notes “not impossible”.
Influenza, meanwhile, spits out hybrid strains all the time – on a much bigger scale known as reassortment. While coronaviruses can also recombine, they are less likely to produce anything “too radically new” in one go, Balloux says, because these spiky pathogens come with their own in-built proofreader – a kind of molecular quality control.
Bauer likens it to the difference between using a cookie cutter to make Christmas biscuits and letting your toddler loose kneading the dough into the right star shape. “That’s probably what allowed it to become larger and more sophisticated than other RNA viruses,” she says.
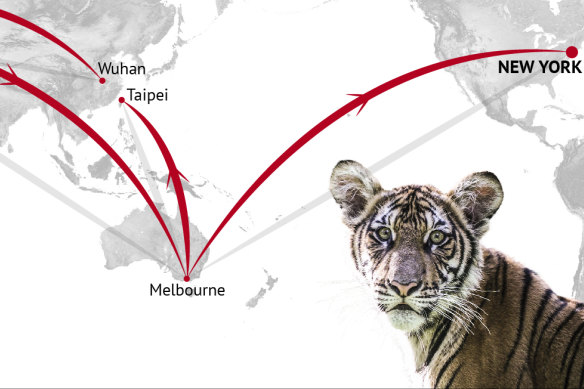
Nadia’s nasal swabs were tested in two different labs, the University of Illinois and Cornell University, then driven through the night to a government lab for further checks – all came back positive.Credit: Stephen Kiprillis, Kirsten Burghard
In Nadia the tiger, one strand of COVID-19 that had stretched all around the world reached the end of the line – the big cat is thought to have caught it from a zookeeper without symptoms in March but did not pass it back to another human.
But she was part of a big cat cluster inside the Bronx Zoo. Nadia was tested first in early April because she seemed the sickest but five tigers and three lions would test positive in total. All are now recovering well, says Professor Karen Terio who helped co-ordinate the tests at her zoological pathology lab at the University of Illinois.
“Nadia was coughing, which is an odd clinical sign for a tiger, so we had a suspicion,” Terio says.
The lab’s virologist, Leyi Wang, who had to devise a whole new lab test for the felines, admits Nadia’s positive still came as a shock. “The zoo had already been shut for weeks when the tiger got sick [at the end] of March … Now we’re looking at the other cats. The tiger’s strain was very similar to the human New York virus but the lion samples look a bit different to the tigers’. It’s not clear yet how they were all infected.”
Balloux notes that the fact the virus can infect other species makes it possible two different lineages – human and animal – could recombine into a hybrid strain down the line. “Some think that’s how we got COVID-19 to begin with. An old bat coronavirus might have mixed with one in a pangolin [a mammal trafficked in China] or maybe it just jumped to the pangolin or [another mammal] that has ACE2 [receptors] like humans and evolved there.”
But he is not worried.
Next to the millions of human COVID-19 cases already confirmed, only a tiny handful of animals, including cats, dogs and ferrets, have tested positive – and then only in mild or asymptomatic cases (or at high deliberate doses in the lab). Aside from a cluster of mink in cramped fur farms in the Netherlands, there has been no sign animals can pass the virus back to people. Experts say this suggests transmission between species (notably humans and their pets) is very weak – and no cause for alarm.
So how did eight big cats fall sick all at once?
The Bronx Zoo did not answer questions on the circumstances or whether staff members had also tested positive, and Terio says that contact tracing work is ongoing and has not been disclosed to her for privacy reasons. While the cluster does offer new clues about what species are particularly susceptible to SARS-CoV-2, she thinks it’s probably “too much of a leap” to say it narrows down the search into the virus’s origins.
Professor Edward Holmes points to another cluster, this time of mink falling severely ill in Dutch farms, as the best place to start looking - mink are also widely farmed in China for their fur. Another lead could lie with the highly endangered and heavily trafficked pangolin which has been found via genomics to carry a similar (though not identical) virus. Holmes says the wildlife trade, including fur farming, remains the most likely origin of the pandemic, as happened with SARS and many other outbreaks.
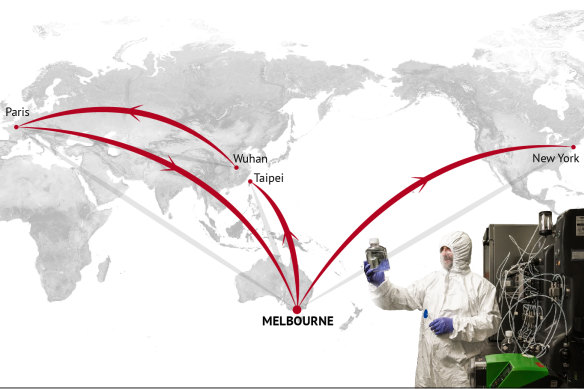
CSIRO scientists are growing both the coronavirus and vaccine candidates in their lab in Geelong, Victoria as part of a global race for a cure.Credit: Stephen Kiprillis, Kirsten Burghard
Fortunately, most scientists say it is unlikely the virus will mutate into something nastier – by its very nature a germ wants to spread, not kill.
Over time Vasan expects it will become milder – the way past pandemic flu strains have as they adapted to their new host. But, also like pandemic flu, COVID could linger on as a seasonal problem that flares up in the cold winter months – at least until it is properly stamped out by vaccination programs.
Back at the CSIRO’s sprawling lab in Geelong, vast stores of the coronavirus – enough for at least two years of vaccine and treatment experimentation – sit in high-security freezers at minus 80 degrees. A virus that mutates slowly is unlikely to lose its bite any time soon. But it's also less likely to change to evade a vaccine.
“I’m not worried it’s going to get worse or mutate out of control of a vaccine,” Balloux says. “It’s already [spreading] out of control now. It's already bad."
Finding an effective vaccine or treatment is critical to getting the world back to some degree of normality while so many of us are still susceptible, Vasan says. Scientists developing vaccines aren’t closing their eyes to the virus genome – much of the work is focused on areas of its molecular structure less likely to mutate.
“It’s like a child in a playground, we’re keeping an eye on it,” Vasan says. “That doesn’t mean we’re worried but we’ll keep looking back while [we work], so we know the child is still there.”
Of course, for all the attention on COVID-19’s genome, he stresses there is still crucial data missing – matching, de-identified clinical records that could show the severity, outcome and personal risk factors of each case.
“Mother Nature is running a giant experiment all around the world right now. That’s where we need to look to see what the mutations mean.”
The problem, Vasan says, is that even very basic patient “metadata” is maddeningly inconsistent across jurisdictions, both interstate and international – and no one is showing much enthusiasm for cleaning it up. “‘Mild’ and ‘moderate’ mean different things everywhere.”
Of the many thousands of virus samples sequenced and shared worldwide, the CSIRO team managed to find just 300 with meaningful, matching patient information.
Their resulting analysis was “underpowered”, Bauer says, but they could already make a preliminary link between changes in clinical outcomes and mutations in the virus’s spike protein. “Imagine what we could do if we had more.”
Only the World Health Organisation could co-ordinate an international approach to streamlining such data into uniform categories, Vasan says, but Australia could still lead the charge with “some show and tell”.
“If we can pull this data together [nationally], we can show the world what it can teach us,” he says. “What we find out could change the game.”
This article was originally published on June 25 and was updated to reflect the second surge of cases in Melbourne.
Get our Morning & Evening Edition newsletters
The most important news, analysis and insights delivered to your inbox at the start and end of each day. Sign up to The Sydney Morning Herald’s newsletter here and The Age’s newsletter here.
Let us explain
If you'd like some expert background on an issue or a news event, drop us a line at explainers@smh.com.au or explainers@theage.com.au. Read more explainers here.